Our First Virtual Organism – E. coli Version 0.1
E. coli is an interesting bacteria with a wide range of roles. It is a tiny rod-shaped lifeform, and completely invisible to the naked eye as it is just 1-2 µM in length. Warm blooded animals including humans depend on E. coli strains that live in the large intestine and prevent harmful bacterial growth while also assisting in the creation of necessary vitamins. Other strains of E. coli are pathogenic, and can cause negative GI symptoms if accidentally consumed. Perhaps the most interesting features of E. coli are those that make it an ideal model organism. E. coli is ideal for many biological experiments as it can adapt to a multitude of environments and reproduce rapidly. We also have several tools available that allow us to manipulate E. coli DNA, making it a perfect organism for experiments in molecular genetics. In this blog we will discuss the motivations of an E. coli cell as well as methods scientists and life programmers will use to interact with this important model organism.
When designing a biosystem, programmers must first consider the motivations of the desired life form. In order to perform any active function, a life form must have the energy to execute the desired behavior. Consequently, metabolism is an integral mechanism, as without it a cell would have no energy available to perform any of the tasks that are necessary for survival. A life programmer might choose to implement metabolic pathways first in order to provide the necessary activation energy for the other various pathways/mechanisms that will need to be modeled. Another inherent component of survivability and organism fitness is reproduction. No designed lifeform would be complete without some description of reproduction. Other behaviors such as motility as described in our previous post, bacterial chemotaxis part 1, will need to be considered and modeled in order to write L++ code which accurately describes the desired lifeform. A life designer will need to account for as many of these basic life functions as possible when they are designing their desired lifeform.
It is important to consider that in the field of biology, there are still many areas we do not understand. Some, such as metabolism, are sufficiently described using reasonable biochemical algorithms. Unfortunately, we do not sufficiently understand the molecular level details of many biological phenomena, and thus are unable to describe the process solely in terms of elementary chemical reactions. For these, we can replace them with mathematical functions with a focus on the design and intention.

Figure 1. Dashboard of a car. Intuitive and clear information visualization of the essential and distilled information for vehicle operation. This image was taken from MACROVECTOR/SHUTTERSTOCK https://www.rd.com/list/car-dashboard-lights-quiz/
Considering the key directives of the organism, how might we go about designing a lifeform like E. coli? Imagine a car dash, the heads-up display (HUD) shows all of the most important information regarding your car’s functioning, and is the primary indicator for when something is going wrong. Similarly to a car’s mechanical functioning, each lifeform has a set of biomechanical functions that must occur properly for the lifeform to survive. A simple interface demonstrating all of these key survival functions creates a very intuitive interface for life programmers designing and debugging an organism.

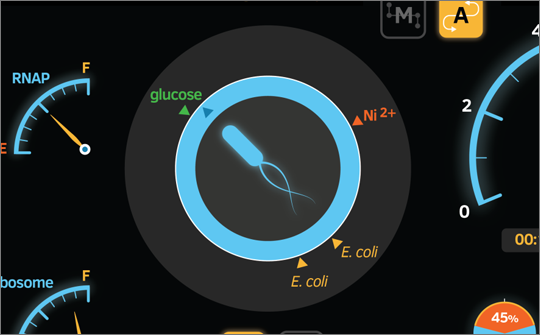
Figure 2. Visualization user interface for E. coli simulation on the life virtualization platform for life programmers. Notice the resemblance to the vehicle dash that we are all familiar with.
Notice the various displays in the above figure, each of the main functions of E. coli are illustrated with a familiar vehicular dash fashion. Key functions are indicated including displays for metabolism, cell replication, navigation, and motility. Many of these have direct analogues with a car’s dash, such as the glucose module being a direct counterpart for the fuel level indicator. Similarly, the navigation module which provides the E. coli cell with directional information is not dissimilar from a compass or even a much more sophisticated navigation system, such as GPS. This simple and easily-operable interface empowers life programmers to very efficiently monitor the life programs they write, enabling quick debugging and fine-tuning of their design.
Let’s take a closer look at some of the other functions displayed on the E. coli dash. There are displays for genetic information processing, as well as replication rate. As previously discussed, a life programmer must consider replication to achieve an accurate model of an organism. Replication requires specialized molecular machineries to form the genetic information, and the displays for RNAP and ribosome provide quick feedback regarding the availability of these genetic components. E. coli cells reproduce asexually via a process known as binary fission. This process entails the replication of DNA, transcription and translation, division of plane determination, and the execution of the physical separation into two identical cell bodies. The indicators for replication rate, division plane, and Z ring constriction give an easy way for programmers to quickly check if the organism is replicating at the desired rate.

Figure 3. Virtual E. coli development roadmap
We are working in collaboration with the Kim Lab at The University of Texas Southwestern Medical Center to write the E. colicode. This code is publicly available and includes many key functions of E. coli. Version 0.1 will include support for the central dogma, cell division, metabolism, and chemotaxis. Full support for DNA replication, transcription into RNA, and translation into proteins is available in this version. Further, cell division can be modeled with DNA replication, midpoint definition by MinCDE system, and FtsZ polymer membrane constriction which simulates the physical separation of the relocated DNA into separate identical cell bodies. Metabolism is also included in this life program including information for glucose storage and metabolization into ATP with the major metabolic pathways of glycolysis, pyruvate oxidation, TCA cycle, and oxidative phosphorylation. Lastly, version 0.1 will include chemotaxis, allowing for simulation of the cell’s motility to respond to increasing glucose levels using biochemical algorithms and flagellar rotation which utilizes a mathematical function.
The Kim Lab and Omphalos Lifesciences Inc will continue to develop and release the virtual E. coli (Figure 3). Version 0.2 will support E. coli populations, growth rates, perturbations (e.g., gene knockout and different media), and so on. Version 1.0 will include over 100 biological processes with some processes described and simulated at high resolution. Some other processes would not be elucidated, given that complete models do not yet exist for them, so these processes are called “unknowns.” Version 1.0 will use “specification” of L++ to describe them – specification will be described later in a different blog post.
Having this virtual E. coli L++ code curated in one location will facilitate understanding with much greater ease and clarity compared to reading numerous dispersed and heterogeneous texts, papers, and figures. The ability to modify and simulate organism code facilitates a much quicker and more thorough understanding of the mechanisms underlying disparate biological processes. Using our simple visualization and simulation interface, life programmers will be able to easily debug their organisms allowing them to efficiently create biochemical pathways, reprogram cells, and design and create organisms.
We expect that numerous researchers around the world will work together to fill in details for unknowns and complete the E. coli code, called version 2.0. We believe that this E. coli code will serve as a springboard for the Human Code Project (HCP), which will be described later in another blog post.
Keep an eye on our website for information regarding the release of E. coli code version 0.1 by the Kim lab. We are excited to share our first virtualized organism, E. coli, on our life virtualization platform, OmVisim. Make sure to stay connected with our blog or social media, eventually, we will virtualize much more complex organisms including humans.




0 Comments