Two abstractions of L++ enables the precise description of biosystems
At first glance any system can look like a complicated web of components; however, it’s important to not get bogged down in every tiny detail. If you were given a recipe, you would only be required to know basic details like which ingredients to use, and how to prepare and heat them. You would not require instructions detailing which knife to use for the job or how to generate the heat needed to prepare your meal. It is critical that systems like these have languages that describe them in order to simplify the complex relationships within each system, and now we are using the L++ programming language to help us simplify the systems of life.
Programming life with L++ requires only that you understand the intended behavior of your molecule of interest to simulate it, the details of your molecule will come on their own. Think about turning on a television. The interface between you and the television is typically a handheld remote with well-defined buttons. You know what each button does and the intended behavior of the signal sent from that button once it is implemented by the television. Even if you do not know how the signal from the remote’s “mute” button is transduced into an electrical signal that is sent wirelessly to the television to turn the sound off, you can still use that button to perform that function. Behaviors of virtual molecules and life forms will be similar to those of the remote by using high-level abstractions coded by L++ to interface with a compiler that implements low-level details about the molecule, cell, or organism of interest. A user could code a simple reaction that is meant to describe the reaction that transforms carbon dioxide into carbonic acid as CO2 -> H2CO3. Although this process can occur on its own slowly, organisms use carbonic anhydrase to drive the reaction to generate carbonic acid faster. Additional parameters like these can be supported by L++ to give more specificity to a program.

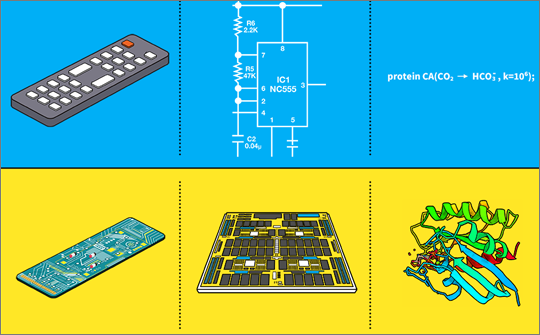
Figure 1: Interface and implementation of various devices and systems. TV viewers turn on TV or switch channels and control volumes via remote controller, without having to instruct how the details of it will be implemented to control the behavior of TV. Similarly, the circuit designer can draw schematics of the circuit, which can be manufactured by the engineers to generate such a circuit board. Using L++ as an interface between biology and programming allows us to design systems that act much like a remote control and a circuit board with short commands we can carry out detailed simulations of biosystems.
While a simple reaction may suffice for some purposes, you may want to expand on the reaction, adding parameters to describe the kinetics of the reaction, enzymes that interact with your molecules of interest, its location within the cell, or environmental conditions that may affect the reaction. All of these parameters can be described in L++ to generate thorough models of the physical properties of molecules and the reactions that they participate in. Let’s take a look at a specific protein: carbonic anhydrase. Using a single line of code we can specify several important details in the interface: The enzyme, its function of converting carbon dioxide to carbonic acid, and the kinetics of the reaction. In using this basal information, the implementation of the code will provide low-level details of the enzymatic activity of carbonic anhydrase such as the zinc binding domain necessary for activation, and the carbon dioxide binding domain that is essential for the production of carbonic acid. The physical properties of molecules that can be computed using the L++ compiler (LCC) also leaves room to create models of larger pathways.

Figure 2: L++ Interface and implementation of carbonic anhydrase. Molecules in the L++ interface can be rigorously described using high level abstractions in code. The L++ compiler (LCC) with databases and structure function prediction algorithms will output detailed information about the target molecule.
The structure of a molecule should be indicative of its function. The molecules within a system should keep a spatial relationship that allows for the proper interactions between each component to carry out the purpose of a system as a whole. Take for example the actin protein that is utilized to form structures that are critical to the structural integrity of a cell. While actin exists as a monomer, it has the ability to bind to itself to form strong filamentous polymers. If we wanted to explore the growth of an actin filament, with just a simple code we could describe the growth of an actin filament from basic monomers to the more complicated helix of a filamentous polymer. The implementation of L++ code into a simulated polymer will cover the low-level details of the polymer such as the spatial relationships between each actin monomer, how the monomers join together, and how the growth of the polymer changes the shape of the filament. With the implementation of L++ code we will be able to describe essentially all biosystems.

Figure 3: L++ Interface and implementation of actin polymerization. High level spatial relationships of molecules in biosystems can be described in L++, and the low level details may be implemented by the L++ compiler (LCC) with databases and capabilities of molecular dynamics simulations.
These two abstractions, descriptions of low-level details of individual molecules and biosystems from high-level language, will allow us to program biosystems at different resolutions. From as small as interactions between individual molecules to spatial organizations of molecules interacting in a biological pathway, to whole-cell interactions, tissue formation, and at a whole organism scale. Refer to our previous blog post Transcending Time, Space, and Resources via Life Virtualization. The different resolutions of life that L++ describes will allow for expedited research and drug development through modeling drug trials on virtual humans which requires accounting for effects on organisms at the molecular, cellular, and organismal level, our first virtual organism, E. coli, is already being built. With requiring only a few lines of code to describe cellular processes, L++ will make molecular modeling more accessible to the scientific community. Keep an eye out for a future blog post focusing on using L++ to generate specifications for living systems.




0 Comments